![]()
MOLECULAR ORGANISATION OF CENTROMERE AND TELOMERE
Centromere
- As earlier described, centromere is a unique region of a chromosome characterized by a constriction, where the two chromatids remain joined together after chromosome replication.
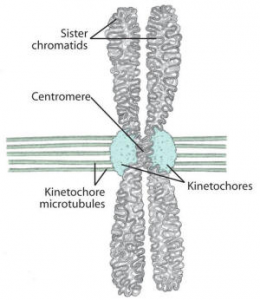
- On each of these centromeres, specialized DNA-protein complexes known as kinetochores assemble.
- Each chromosome has two kinetochores, one on each sister chromatid, facing in opposite directions.
- These kinetochores are the sites for binding of kinetochore microtubules (spindle), which assist in the movement of chromosomes during anaphase.
- The DNA sequence representing the centromere contain the information specifying the assembly of kinetochore.
- These centromeric DNA sequences have been isolated and characterized in a number of organisms but, unlike the telomeric sequences, have not been found to be conserved.
- The kinetochore functions as the “microtubule organizing Centre” (MTOC) of the chromosome. Such centrios also occur at the poles, in the vicinity of centriole.
- A single microtubule is sufficient to move a chromosome. but, the number of microtubules attached to a single chromosome .
- In yeast there is only one microtubule per kinetochore, but in human, 20 to 40 microtubules are found attached to one kinetochore.
- The centromere is surrounded by heterochromatin that can be observed by the C-banding technique. Satellite DNA sequences are located on both the sides of centromeres.
- The minimum region of a chromosome necessary for centromeric function is called a CEN fragment or CEN region.
- For example – In case of yeast, Each of the 17 chromosomes of yeast contains a different centromeric sequence, each sequence being with in 120 bp in length.
- However, all the sequences contain substantial regions of homology, and can be inverted or swapped from one chromosome to another without loss of function.
- The yeast centromeric DNA contains three distinguishable sequences.
(i) Conserved element I (CDE I):
- It is composed of 9 bp and is located at the left end of the centromere; it shows minor variations.
(ii) Conserved element II (CDE II):
- This element is the middle region containing 80-90 bp. A=T rich sequences constitute more than 90% of this region.
(iii) Conserved element III (CDE III):
- This element consists of 11 bp and is located at the right end of the centromere (Fig. 8.8). It is a highly conserved sequence.
- Point mutations in the CCG sequence of the conserved element III leads to an inactivation of the centromeric function.
- Mutations in the other elements, however, cause only a reduction in the centromeric function.
- Certain proteins or protein complexes bind to specific regions of the CEN sequences.

- Protein CBF-I binds to CDEI element. Proteins CBF-IIIA, CBF-IIIB and CBF-IIIC from a complex (M.W. 2.4 × 105 Daltons) which binds to the CDE-III region.
- This protein complex has some motor activity due to which the centromeric region of the chromosome becomes attached to microtubules.
- Mitotic chromosome movement is inhibited when mutation occurs in the genes coding for CBF-III proteins.
- The yeast centromeric sequences bind to specific proteins which initiate to form a kinetochore (multi-protein complex).
- The kinetochore, in turn, binds to the end of a single microtubule. Mammalian centromeres are thought to contain different and much longer DNA sequences.
- They form larger kinetochores each of which binds to several microtubules.
Telomere
- The sequences at the ends of eukaryotic chromosomes, called telomeres, play critical roles in chromosome replication and maintenance.
- Telomeres were initially recognized as distinct structures because broken chromosomes were highly unstable in eukaryotic cells, implying that specific sequences are required at normal chromosomal termini.

- In recent years the structure of telomeres in a wide variety of organisms has been studied to demonstrate that telomeres are highly conserved elements throughout the eukaryotes, both in structure and function.
- Telomeric DNA has been shown to consist of simple randomly repeated sequences, characterized by clusters of G residues in one strand and C residues in the other.
- Another feature is a 3’ overhang (12-16 nucleotides in length) of the G-rich strand. For example, the sequence of telomere repeats in humans and other mammals is AGGGTT, and the telomere repeat in Tetrahymena is GGGGTT.
- These sequences are repeated hundreds or thousands of times, thus spanning up to several kilobases, and terminate with an overhang of single-stranded DNA.
- Recent results suggest that the repeated sequences of telomere DNA form loops at the ends of chromosomes, thereby protecting the chromosome termini from degradation
- Telomeres play a critical role in replication of the ends of linear DNA molecules.
- The same repeated sequence is found at the ends of all chromosomes in a species and the same telomere sequence may occur in widely divergent species, such as humans, some acellular slime molds (trypanosome) and the fungi like Neurospora.
- At every telomere, as much as 10 kilobase of this repeat sequence may occur.
- The telomric DNA is also complexed with non-histone proteins,the complex structure being associated with nuclear lamina, as shown in Oxytricha, a ciliated protozoan.
- The telomeric DNA is synthesized under the influence of telomerase.
- An enzyme which has been shown to be a ribonucleoprotien, whose RNA component works as a template for synthesis of telomeric DNA repeats and protein component has reverse transcriptase (RT) activity.
- Recently (In 1997), proteins have been isolated from telomerase in yeast and a ciliated protozoan (Euplotes) which could be the RT component of telomerase.
- These were p123 in Euplotes and Est2 in yeast. In recent years, efforts have also been made to understand, how the telomere length is monitored in telomerase, because uncontrolled activity of telomerase will lead to indefinite elongation of telomeres.
- Proteins binding to telomere repeats have been identified, which block the elongation of telomeres.
- These include rap1p in budding yeast, Taz1p in fission yeast and TRF in humans.
- The significance of these proteins lies in the following two observations made in recent years:
- (i) Shortening of telomere tract (repeat sequence) is associated with senescence and aging;
- (ii) Controlled elongation of telomere is an essential step in tumour formation and oncogenesis.
- However, even in tumour cells, this telomere length does not increase in an uncontrolled manner, and the elongation is regulated.
- However, we still do not know, ‘how does the cell measure the telomere length, and how is this information used to regulate the length of telomeres’?













