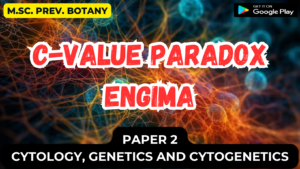
- In many repressible operons, transcription that initiates at the promoter can terminate prematurely in a leader region that precedes the first structural gene. (i.e. the polymerase terminates transcription before it gets to the first gene in the operon).
- This phenomenon is called ATTENUATION; the premature termination of transcription.
- Although attenuation is seen in a number of operons, the mechanism is best understood in those repressible operons involved in amino acid biosynthesis.
- In these instances attenuation is regulated by the availability of the cognate aminoacylated t-RNA.
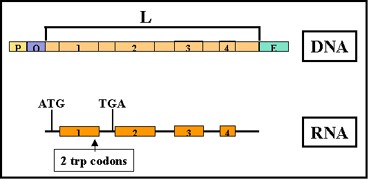
Figure 2.39 : Mechanism of attenuation
- Mechanism (See Figure 2.39) – When transcription is initiated at the promoter,
- it actually starts before the first structural gene and a leader transcript is made.
- This leader region contains a start and a stop signal for protein synthesis.
- Since bacteria do not have a nuclear membrane, transcription and translation can occur simultaneously.
- Thus, a short peptide can be made while the RNA polymerase is transcribing the leader region.
- The test peptide contains several tryptophan residues in the middle of the peptide.
- Thus, if there is a sufficient amount of tryptophanyl-t-RNA to translate that test peptide, the entire peptide will be made and the ribosome will reach the stop signal.
- If, on the other hand, there is not enough tryptophanyl-t-RNA to translate the peptide, the ribosome will be arrested at the two tryptophan codons before it gets to the stop signal.
- The sequence in the leader m-RNA contains four regions, which have complementary sequences (Figure 2.40).

Figure 2.40 : Formation of stem-loops
- Thus, several different secondary stem and loop structures can be formed.
- Region 1 can only form base pairs with region 2; region 2 can form base pairs with either region 1 or 3; region 3 can form base pairs with region 2 or 4; and region 4 can only form base pairs with region 3.
- Thus three possible stem/loop structures can be formed in the RNA.
- region 1:region 2
- region 2:region 3
- region 3:region 4
- One of the possible structures (region 3 base pairing with region 4) generates a signal for RNA polymerase to terminate transcription (i.e. to attenuate transcription).
- However, the formation of one stem and loop structure can preclude the formation of others.
- If region 2 forms base pairs with region 1 it is not available to base pair with region 3.
- Similarly if region 3 forms base pairs with region 2 it is not available to base pair with region 4.
- The ability of the ribosomes to translate the test peptide will affect the formation of the various stem and loop structures Figure 2.41.
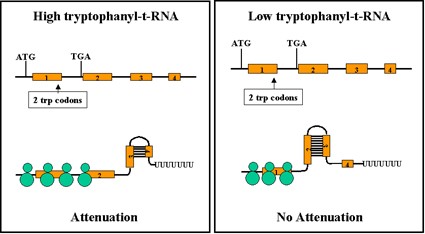
Figure 2.41 : Mechanism of atteunation
- If the ribosome reaches the stop signal for translation it will be covering up region 2 and thus region 2 will not available for forming base pairs with other regions.
- This allows the generation of the transcription termination signal because region 3 will be available to pair with region 4.
- Thus, when there is enough tryptophanyl-t-RNA to translate the test peptide attenuation will occur and the structural genes will not be transcribed.
- In contrast, when there is an insufficient amount of tryptophanyl-t-RNA to translate the test peptide no attenuation will occur.
- This is because the ribosome will stop at the two tryptophan codons in region 1, thereby allowing region 2 to base pair with region 3 and preventing the formation of the attenuation signal (i.e. region 3 base paired with region 4).
- Thus, the structural genes will be transcribed.
(2) In Viruses
- Viruses consist of a nucleic acid (DNA or RNA) enclosed in a protein coat (known as a capsid).
- The capsid may be a single protein repeated over and over, as in tobacco mosaic virus (TMV).
- It may also be several different proteins, as in the T-even bacteriophages.
- Once inside the cell, the nucleic acid follows one of two paths: lytic or lysogenic.
- Retroviruses, such as Human Immuno-difficiency Virus (HIV), also include the enzyme reverse transcriptase with the viral RNA.
- Reverse transcriptase makes a single-stranded viral DNA copy of the single-stranded viral RNA.
- The single stranded viral DNA is subsequently turned into a double-stranded DNA.
- The lytic cycle occurs when the viral DNA immediately takes over the host cell (remember that viruses are obligate intracellular parasites) and begins making new viruses.
- Eventually the new viruses cause the rupture (or lysis) of the cell, releasing those new viruses to continue the infection cycle.
- The lysogenic cycle occurs when the viral DNA is incorporated into the host DNA as a prophage.
- When the cell replicates the prophage is passed along as if it were host DNA.
- Sometimes the prophage can emerge from the host chromosome and enter the lytic cycle spontaneously once every 10,000 cell divisions.
- Ultraviolet light and x-rays may also trigger emergence of the prophage.
- Transduction is the transfer of host DNA from one cell to another by a virus (Figure 2.42).
- Some bacteriophages are temperate since they tend to go lysogenic rather than lytic.
- These types of viruses are able to transduce fragments of the host DNA.
fig 2.42a
fig 2.42.b
- Transposons are DNA fragments incorporated into the chromosomal DNA (Figure 2.43). Unlike episomes and prophages, transposons contain a gene producing an enzyme that catalyzes insertion of the transposon at a new site.
- They also have repeated sequences 20-40 nucleotides in length at each end.
- Insertion sequences are short (600-1500 base pairs long) simple transposons that do not carry genes beyond those essential for insertion of the transposon into E. coli. Complex transposons are much larger and carry additional genes.
- Genes incorporated in a complex transposon are known as jumping genes since they can move about on the chromosome (even from chromosome to chromosome).
- Often the complex transposons are flanked by simple transposons

Figure 2.43 : Trans-posons and their relationship to other genes.






