RNA Splicing
- The other major type of modification that takes place in eukaryotic pre-mRNA is the removal of introns by RNA splicing.
- This occurs in the nucleus following transcription but before the RNA moves to the cytoplasm.
- Consensus sequences and the spliceosome Splicing requires the presence of three sequences in the intron.
- One end of the intron is referred to as the 5′ splice site, and the other end is the 3′ splice site (Figure 2.25); these splice sites possess short consensus sequences.
- Most introns in pre-mRNA begin with GU and end with AG, suggesting that these sequences play a crucial role in splicing.
- Changing a single nucleotide at either of these sites does indeed prevent splicing.
- A few introns in pre-mRNA begin with AU and end with AC.
- These introns are spliced by a process that is similar to that seen in GU. . . AG introns but utilizes a different set of splicing factors.
- This discussion will focus on splicing of the more common GU. . . AG introns.
Table 2.1: RNA–RNA interactions in pre-mRNA splicing
| Interaction | Function |
| U1 with 5′ splice site | U1 attaches to 5′ end of intron; commits intron to splicing; no direct role in splicing |
| U2 with branch point | Positions 5′ end of intron near branch point for lariat formation |
| U2 with U6 | Holds 5′ end of intron near branch point |
| U6 with 5′ splice site | Positions 5′ end of intron near branch point |
| U5 with 3′ end of first exon | Anchors first exon to spliceosome subsequent to cleavage; juxtaposes two ends of exon for splicing |
| U5 with 3′ end of one exon and 5′ end of the other | Juxtaposes two ends of exon for splicing |
| U4 with U6 | Delivers U6 to intron; no direct role in splicing |
- The third sequence important for splicing is at the so-called branch point, which is an adenine nucleotide that lies from 18 to 40 nucleotides upstream of the 3′ splice site (Figure 2.25).
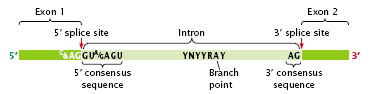
Figure 2.25: Splicing of pre-mRNA requires consensus sequences.
- In the consensus sequence surrounding the branch point (YNYYRAY) Y is any pyrimidine, R is any purine, A is adenine, and N is any base.
- The sequence surrounding the branch point does not have a strong consensus but usually takes the form YNYYRAY (Y is any pyrimidine, N is any base, R is any purine, and A is adenine).
- The deletion or mutation of the adenine nucleotide at the branch point prevents splicing.
- Splicing takes place within a large complex called the spliceosome, which consists of several RNA molecules and many proteins.
- The RNA components are small nuclear RNAs; these snRNAs associate with proteins to form small ribonucleoprotein particles.
- Each snRNP contains a single snRNA molecule and multiple proteins.
- The spliceosome is composed of five snRNPs, named for the snRNAs that they contain (U1, U2, U4, U5, and U6), and some proteins not associated with an snRNA.
- The process of splicing To illustrate the process of RNA splicing, we’ll first consider the chemical reactions that take place.
- Then we’ll see how these splicing reactions constitute a set of coordinated processes within the context of the spliceosome.
- Before splicing takes place, an upstream exon (exon 1) and a downstream exon (exon 2) are separated by an intron (Figure 2.26).
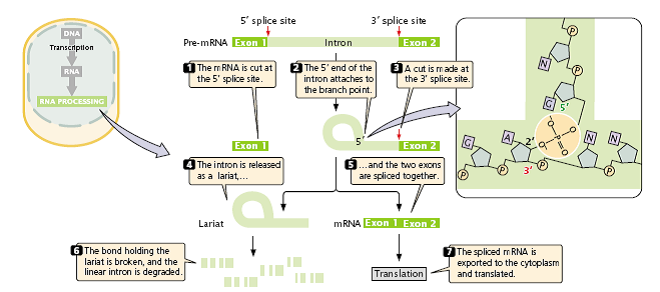
Figure 2.26: The splicing of nuclear introns requires a two-step process.
- First, cleavage takes place at the 5′ splice site, and a lariat is formed by the attachment of the 5′ end of the intron to the branch point.
- Second cleavage takes place at the 3′ splice site, and two exons are spliced together.
Pre-mRNA is spliced in two distinct steps.
- In the first step, the pre-mRNA is cut at the 5′ splice site.
- This cut frees exon 1 from the intron, and the 5′ end of the intron attaches to the branch point; that is, the intron folds back on itself, forming a structure called a lariat.
- The guanine nucleotide in the consensus sequence at the 5′ splice site, bonds with the adenine nucleotide, at the branch point.
- This bonding is accomplished through transesterification, a chemical reaction in which the OH group on the 2′-carbon atom of the adenine nucleotide at the branch point attacks the 5′ phosphodiester bond of the guanine nucleotide at the 5′ splice site, cleaving it and forming a new 5’–2′ phosphodiester bond between the guanine and adenine nucleotides.
- In the second step of RNA splicing, a cut is made at the 3′ splice site and, simultaneously, the 3′ end of exon 1 becomes covalently attached (spliced) to the 5′ end of exon 2.
- This bond also forms through a transesterification reaction, in which the 3′-OH group attached to the end of exon 1 attacks the phosphodiester bond at the 3′ splice site, cleaving it and forming a new phosphodiester bond between the 3′ end of exon 1 and the 5′ end of exon 2; the intron is released as a lariat.
- The intron becomes linear when the bond breaks at the branch point and is then rapidly degraded by nuclear enzymes.
- The mature mRNA consisting of the exons spliced together is exported to the cytoplasm where it is translated.
- Although splicing is illustrated in Figure 2.27 as a two-step process, the reactions are in fact coordinated within the spliceosome.
- A key feature of the spliceosome is a series of interactions between the mRNA and snRNAs and between different snRNAs (summarized in Table 2.1).
- These interactions depend on complementary base pairing between the different RNA molecules and bring the essential components of the pre-mRNA transcript and the spliceosome close together, which makes splicing possible.
- The spliceosome is assembled on the pre-mRNA transcript in a step-by-step fashion (Figure 2.27).
- First, snRNP U1 attaches to the 5′ splice site, and then U2 attaches to the branch point.
- A complex consisting of U5 and U4–U6 (which form a single snRNP) joins the spliceosome.

Figure 2.27: RNA splicing takes place within the spliceosome
- At this point, the intron loops over and the 5′ splice site is brought close to the branch point. U1 and U4 disassociate from the spliceosome.
- The 5′ splice site, 3′ splice site, and branch point are in close proximity, held together by the spliceosome.
- The two transesterification reactions take place, joining the two exons together and releasing the intron as a lariat.
- An animation of the splicing process nuclear organization RNA splicing takes place in the nucleus and must occur before the RNA can move into the cytoplasm.
- For many years, the nucleus was viewed as a biochemical soup, in which components such as the spliceosome diffused and reacted randomly.
- Now, the nucleus is believed to have a highly ordered internal structure, with transcription and RNA processing taking place at particular locations within it.
- By attaching fluorescent tags to pre-mRNA and using special imaging techniques, researchers have been able to observe the location of pre-mRNA as it is transcribed and processed.
- The results of these studies revealed that intron removal and other processing reactions take place at the same sites as those of transcription (Figure 2.28), suggesting that these processes may be physically coupled.
- This suggestion is supported by the observation that part of RNA polymerase II is also required for the splicing and 3′ processing of pre-mRNA.
Figure 2.28: Group I introns undergo self-splicing.
(a) Secondary structure of a group I intron. (b) Self-splicing of a group I intron
Self-Splicing Introns –
- Some introns are self-splicing, meaning that they possess the ability to remove themselves from an RNA molecule.
- These self-splicing introns fall into two major categories.
- Group I introns are found in a variety of genes, including some rRNA genes in protists, some mitochondrial genes in fungi, and even some bacteriophage genes.
- Although the lengths of group I introns vary, all of them fold into a common secondary structure with nine looped stems (Figure 2.28), which are necessary for splicing.
- Transesterification reactions are required for the splicing of group I introns (Figure 2.29).
Figure 2.29: Group II introns undergo self-splicing by a different mechanism from that for group I introns.
(a) Secondary structure of a group II intron.
(b) Self-splicing of group II introns, which is similar to the splicing of nuclear introns.
Alternative Processing Pathways –
- Another finding that complicates the view of a gene as a sequence of nucleotides that specifies the amino acid sequence of a protein is the existence of alternative processing pathways, in which a single pre-mRNA is processed in different ways to produce alternative types of mRNA, resulting in the production of different proteins from the same DNA sequence.
- One type of alternative processing is alternative splicing, in which the same pre-mRNA can be spliced in more than one way to yield multiple mRNAs that are translated into proteins with different amino acid sequences (Figure 2.30a).
- Another type of alternative processing requires the use of multiple 3′ cleavage sites (Figure 2.30b); two or more potential sites for cleavage and polyadenylation are present in the pre-mRNA.
Figure 2.30: Eukaryotic cells have alternative pathways for processing pre-mRNA.
(a) With alternative splicing; pre-mRNA can be spliced in different ways to produce different mRNAs.
(b) With multiple 39 cleavage sites, there are two or more potential sites for cleavage and polyadenylation; use of the different sites produces mRNAs of different lengths
- In our example, cleavage at the first site produces a relatively short mRNA, compared with the mRNAs produced through cleavage at other sites.
- Both alternative splicing and multiple 3′ cleavage sites can exist in the same pre-mRNA transcript; an example is seen in the mammalian calcitonin gene, which contains six exons and five introns (Figure 2.31a).
- The entire gene is transcribed into pre-mRNA (Figure 2.31b).
- There are two possible 3′ cleavage sites. In cells of the thyroid gland, 3′ cleavage and polyadenylation take place after the fourth exon, and the first three introns are then removed to produce a mature mRNA consisting of exons 1, 2, 3, and 4 (Figure 2.31c). This mRNA is translated into the hormone calcitonin.
- In brain cells, the identical pre-RNA is transcribed from DNA, but it is processed differently. Cleavage and polyadenylation take place after the sixth exon, yielding an initial transcript that includes all six exons.
- During splicing, exon 4 (part of the calcitonin mRNA) is removed, along with all the introns; so only exons 1, 2, 3, 5, and 6 are present in the mature mRNA (Figure 2.31d).
- When translated, this mRNA produces a protein called calcitonin-gene-related peptide (CGRP), which has an amino acid sequence quite different from that of calcitonin.
Figure 2.31: Pre-mRNA encoded by the calcitonin gene undergoes alternative processing
- Alternative splicing may produce different combinations of exons in the mRNA, but the order of the exons is not usually changed.
- Different processing pathways contribute to gene regulation.






