Molecular mechanisms of Recombination
Molecular mechanisms of Recombination
- DNA replication with 100% fidelity is a nice feature to keep offspring in just the genetic background of the species.
- But to get there, or to evolve further, requires genetical changes, one of which results from recombination of (near) homologous parts of DNA.
- The nature of structural changes in DNA neccessary to result in homologous genetic recombination were layed out by R. HOLLIDAY in 1964, and in subsequent years the crossover-structures were visualized by electron microscopy (Figure 3.3).
Figure 3.3: A ‘X’-form that has been prepared for the electron microscope in the presence of a high concentration of formamide.
- Under these conditions the DNA double helix is stressed, and those regions particulary rich in AT base pairs undergo a localized denaturation.
- This sequence-specific denaturation allows the homologous arms in the molecule to be identified.
- Furthermore, the covalent strand connections in the region of the crossover can be seen. In this and other 80 open molecules, the homologous arms are in a trans configuration (Photo: H. POTTER und D. DRESSLER, 1976).
- The actual conformation of a DNA crossover was speculated to be a four-way-junction with separate DNA helices, or with stacked helices in either a parallel or an antiparallel orientation of the helices.
- The models had to allow for branch migration, else no exchange of genetic material would happen.
Breakage – Fusion (Reunion) – Bridge Cycle, Control Elements And Unstable Genes –
- Since the beginning of the century has it been known that unstable or variable gene loci occur in plants although the drastically enhanced mutability and the increased back mutation rate could not be explained at first.
- The decisive breakthrough was accomplished by B. McCLINTOCK with her studies on maize chromosomes published in 1947 and 1951.
- The basis of these were her earlier observations and analyses (1938) on breakage-fusion (reunion)-bridges.
- Their occurrence could be correlated with the restructuring in chromosomes.
- Bridges are formed during anaphase whenever two chromosomes fuse at their ends generating a fusion product with two centromers.
- If these two are subsequently torn to different poles then will inevitably occur a chromosomal fraction.
- During the following S-phase of the interphase nucleus is a chromatid with a fraction at its terminus replicated in just the same way as the other chromosomes leading again to a fusion of the homologous chromatids.
- Consequently can a chromosome consisting of just one chromatid but two centromers be found in the subsequent mitosis instead of a chromosome out of two chromatids and one centromer.
- The consequence is a second fraction during anaphase where the second round of the cycle starts.
- B. McCLINTOCK recognized that the fraction cannot occur at any site of the chromosome but is restricted to certain sections that she called Ds (dissociation).
- These were obviously DNA segments contributing to the formation of translocations, deletions, inversions and to the generation of ring-shaped chromosomes.
- The first fraction causes similar fractions in the mitosis cycles of following generations.
- They happen during ontogenesis at different times and sites.
- The segment Ds, a mutator gene, behaves like a multiple allele (or, even better, like a pseudoallele) that can be located at different gene loci. It may also vary in structure.
- This mutator can insert itself into other genes thus rendering them inactive.
- It is a control element that changes its place within the chromosome, jumping or wandering around and causing mutations wherever it inserts (the mutators are also called jumping genes).
- It soon became clear that a further set of elements has to exist: the Ac (activation) elements.
- A chromosomal fraction or a translocation of a Ds element has to be supported by an Ac element. An Ac element can also be regarded as a multiple allele.
- It may occur at the most different sites in all chromosomes.
- To analyze its effect further concentrated B. McCLINTOCK on the study of genes that determine the colour of maize grains.
- One of the most important is the C-locus that causes a dark red staining of the aleuron layer and the pericarp of the maize grain in a dominant condition.
- If a Ds-element jumps into the gene, colour synthesis is interrupted and colourless (yellow) grains result.
- An Ac activity within these grains causes a pattern of dark red areas on a light ground.
- This is explained by a reestablishment of the old state since the Ac element removes the Ds from the C-locus.
- This happens in several cells during the development of the maize grain.
- These cells are the origin of the aleuron layer and the pericarp and the back mutation can only be perceived in the clones that form out of the changed cells.
- Today are a number of gene loci known that can be influenced by the Ds-Ac-system or other control elements.
- The detection of the spm-system (suppressor-mutator) and the elucidation of its function showed that the control elements do not only act as switches (a yes/ no decision) but that they do modulate the degree of gene expression, too.
- The genetic analyses of B. McCLINTOCK were not understood for years.
- Only when insertion elements and transposons were found in bacterial DNA during the late sixties did an analogy between them and the control elements show up.
- These genetic data fitted neatly with molecular biological models (B. NEVERS and H. SAEDLER, 1977, H-P. DÖRING and P. STARLINGER, 1984).
- Mrs BARBARA McCLINTOCK was awarded the Nobel prize for medicine and physiology for her pioneer achievements. P.NEVERS, N. S. SHEPHERD and H. SAEDLER listed the ‘unstable plant genes’ described in literature at the beginning of 1986.
- It shows that such genes have been found in more than 30 species.
- Many of the respective mutants with names like variegate, marmorata, maculata or variabilis are on the market as ornamental plants due to their irregularly spotted flowers or leaves.
Holliday junction: central intermediate of genetic recombination –
- DNA replication with 100% fidelity is a nice feature to keep offspring in just the genetic background of the species.
- But to get there, or to evolve further, requires genetic changes, one of which results from recombination of (near) homologous parts of DNA.
- The nature of structural changes in DNA neccessary to result in homologous genetic recombination were layed out by Holliday in 1964, and in subsequent years the crossover-structures were visualized by electron microscopy.
- The actual conformation of a DNA crossover was speculated to be a four-way-junction with separate DNA helices, or with stacked helices in either a parallel or an antiparallel orientation of the helices.
- The models had to allow for branch migration, else no exchange of genetic material would happen.
- During branch migration hydrogen bonds between paired bases have to be broken and others reformed instead.
- On average the energy for braking and reforming these bonds will cancel each other – but in real existing DNA not all base pairs are created equal.
- This calls for the action of enzymes to overcome the neccessary activation energy. And enzymes are needed anyway to resolve the four-way-junctions into separate helices.
- In E. coli. e.g. there exists an enzyme system (RuvABC) the components of which hold the Holliday junction (RuvA), swivel the DNA strands to enable branch migration (RuvB) and finally cut the junction (RuvC). A DNA ligase restores intact double helices.

Figure 3.4: RecA protein-dsDNA complex imaged by atomic force microscopy (AFM):
- Homologous genetic recombination is a highly dynamic process, in contrast to X-ray crystallography relaying on static structures.
- So it took to the end of the previous millenium to get an atomic detail view of relevant structures.
- You may see here the structure of a four-way Holliday-junction formed by homologous DNA strands, a RuvA-tetramer complexed to a static Holliday-junction, the motor driving branch migration, and a Holliday-junction resolving enzyme.
- The Ruv-System of E. coli is in itself a dynamic complex.
- During branch migration two tetramers of RuvA hover on both sides of a cruciform DNA, with multimeric RuvB clamping two of the DNA strands to wind them.
- This complex is not accessible for RuvC. In order for the resolvase to act, one of the RuvA tetramers has to be dissociated so that one side of the DNA junction is amenable to strand separation.
- In vitro the tetramer-octamer-equilibrium is subject to the salt concentration of the buffer. Conditions neccessary for crystallisation of the complex resulted in tetrameric RuvA complexed to the DNA.
- Our research is focused on the molecular mechanisms of genetic recombination, with the long-term objective being the reconstitution of in vitro systems that accurately reproduce the cellular processes.
- We are characterizing the biochemical properties of proteins essential to homologous recombination, in prokaryotes, eukaryotes, and Archaea.
- In E. coli, the RecA, RecBCD, RecQ, RuvABC, and SSB proteins, and a specific DNA sequence called Chi, are essential to homologous recombination.
- The RecA protein possesses the unique ability to pair homologous DNA molecules (Figure 3.4) and to promote the subsequent exchange of DNA strands.
- Since RecA protein is the prototypic DNA strand exchange protein, we are interested in the biochemical mechanism of protein-mediated recognition and exchange of homologous DNA strands.
- The RecBCD enzyme is both a DNA helicase and a nuclease with the remarkable properties that its nuclease activity, but not its helicase activity, is attenuated by interaction with the Chi sequence, and that it will actively load RecA protein onto ssDNA.
- RecQ protein is a helicase that can also effect recombination events. SSB protein is an ssDNA binding protein that stimulates the activities of RecA, RecBCD, and RecQ proteins by virtue of its ability to bind ssDNA.
- Recently, we reconstituted an in vitro pairing reaction that requires the concerted action of each of these proteins; the role of each protein in this reaction is under investigation.
- We are also studying the biochemistry of homologous recombination in the yeast, S. cerevisiae and the archaeon, S. solfataricus. Rad51 and RadA proteins are the RecA protein homologues, respectively.
- In yeast, at least three ancillary proteins are needed for Rad51 protein-mediated DNA strand exchange: these include the RP-A, Rad52, and Rad54 proteins.
- We are studying the mechanism of these reconstituted reactions
- General Recombination Is Guided by Base-pairing Interactions Between Complementary Strands of Two Homologous DNA Molecules – General recombination involves DNA strand-exchange intermediates that require some effort to understand.
- Although the exact pathway followed is likely to be different in different organisms, detailed genetic analyses of viruses, bacteria, and fungi suggest that the major outcome of general recombination is always the same.
(1) Two homologous DNA molecules “cross over”; that is, their double helices break and the two broken ends join to their opposite partners to re-form two intact double helices, each composed of parts of the two initial DNA molecules (Figure 3.5).
(2) The site of exchange (that is, where a red double helix is joined to a green double helix (in Figure 3.5) can occur anywhere in the homologous nucleotide sequences of the two participating DNA molecules.
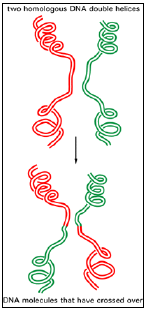
Figure 3.5: General recombination.
- The breaking and rejoining of two homologous DNA double helices creates two DNA molecules that have “crossed over.”
(3) At the site of exchange, a strand of one DNA molecule becomes base-paired to a strand of the second DNA molecule to create a staggered joint (usually called a heteroduplex joint) between the two double helices (Figure 3.6). The hetero-duplex region can be thousands of base pairs long; we shall explain later how it forms.
(4) No nucleotide sequences are altered at the site of exchange; the cleavage and re-joining events occur so precisely that not a single nucleotide is lost or gained. Despite this precision, general recombination creates DNA molecules of novel sequ-ence: the heteroduplex joint can contain a small number of mismatched base pairs, and, more important, the two DNAs that cross over are usually not exactly the same on either side of the joint.
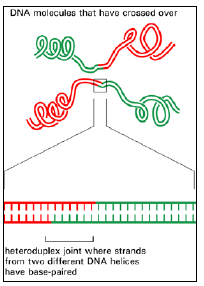
Figure 3.6: A heteroduplex joint.
- This structure unites two DNA molecules where they have crossed over. Such a joint is often thousands of nucleotides long
- The mechanism of general recombination ensures that two regions of DNA double helix undergo an exchange reaction only if they have extensive sequence homology.
- The formation of a heteroduplex joint requires that such homology be present because it involves a long region of complementary base-pairing between a strand from one of the two original double helices and a complementary strand from the other.
- But how does this heteroduplex joint arise, and how do the two homologous regions of DNA at the site of crossing-over recognize each other? As we shall see, recognition takes place by means of a direct base-pairing interaction.
- The formation of base pairs between complementary strands from the two DNA molecules then guides the general recombination process, allowing it to occur only between long regions of matching DNA sequence.
General Recombination Can Be Initiated at a Nick in One Strand of a DNA Double Helix –
- Each of the two strands in a DNA molecule is helically wound around the other.
- As a result, extensive base-pair interactions can occur between two homologous DNA double helices only if a nick is first made in a strand of one of them, freeing that strand for the unwinding and rewinding events required to form a heteroduplex with another DNA molecule.
- For the same reason, any exchange of strands between two DNA double helices requires at least two nicks, one in a strand of each interacting double helix. Finally, to produce the heteroduplex joint illustrated in Figure 3.6, each of the four strands present must be cut to allow each to be joined to a different partner.
- In general recombination, these nicking and resealing events are coordinated so that they occur only when wo DNA helices share an extensive region of matching DNA sequence.
- There is evidence from a number of sources that a single nick in only one strand of a DNA molecule is sufficient to initiate general recombination.
- Chemical agents or types of irradiation that introduce single strand nicks, for example, will trigger a genetic recombination event.
- Moreover, one of the special proteins required for general recombination in E. coli the RecBCD protein has been shown to make single strand nicks in DNA molecules.
- The RecBCD protein is also a DNA helicase, hydrolyzing ATP and traveling along a DNA helix transiently exposing its strands.
- By combining its nuclease and helicase activities, the RecBCD protein will create a single-stranded “whisker” on the DNA double helix (Figure 3.7). Figure 3.8 shows how such a whisker could initiate a base-pairing interaction between two complementary stretches of DNA double helix.
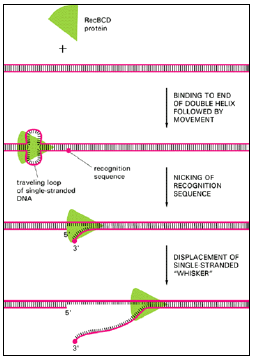
Figure 3.7: One way to start a recombination event.
- The RecBCD protein is an enzyme required for general genetic recombination in E. coli.
- The protein enters the DNA from one end of the double helix and then uses energy derived from the hydrolysis of bound ATP molecules to propel itself in one direction along the DNA at a rate of about 300 nucleotides per second.
- A special recognition site (a DNA sequence of eight nucleotides scattered throughout the E. coli chromosome) is cut in the traveling loop of DNA created by the RecBCD protein, and thereafter a single-stranded whisker is displaced from the helix, as shown.
- This whisker is thought to initiate genetic recombination by pairing with a homologous helix, as in Figure 3.8.
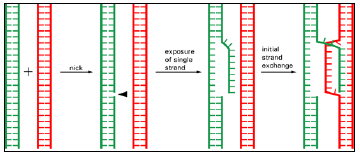
Figure 3.8: The initial strand exchange in general recombination.
- A nick in a single DNA strand frees the strand, which then invades a homologous DNA double helix to form a short pairing region with one of the strands in the second helix.
- Only two DNA molecules that are complementary in nucleotide sequence can base-pair in this way and thereby initiate a general recombination event.
- All of the steps shown here can be catalyzed by known enzymes (see Figures 3.7 and 3.11).
- DNA Hybridization Reactions Provide a Simple Model for the Base-pairing Step in General Recombination – In its simplest form, the type of base-pairing interaction central to general recombination can be mimicked in a test tube by allowing a DNA double helix to re-form from its separated single strands.
- This process, called DNA renaturation or hybridization, occurs when a rare random collision juxtaposes complementary nucleotide sequences on two matching DNA single strands, allowing the formation of a short stretch of double helix between them.
- This relatively slow helix nucleation step is followed by a very rapid “zippering” step as the region of double helix is extended to maximize the number of base-pairing interactions (Figure 3.9).
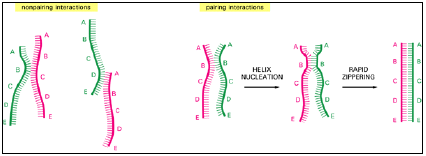
Figure 3.9: DNA hybridization.
- DNA double helices re-form from their separated strands in a reaction that depends on the random collision of two complementary strands. Most such collisions are not productive, as shown at the left, but a few result in a short region where complementary base pairs have formed (helix nucleation).
- A rapid zippering then leads to the formation of a complete double helix.
- A DNA strand can use this trial-and-error process to find its complementary partner in the midst of millions of non-matching DNA strands.
- Trial-and-error recognition of a complementary partner DNA sequence appears to initiate all general recombination events.
- Formation of a new double helix in this way requires that the annealing strands be in an open, unfolded conformation.
- For this reason in vitro hybridization reactions are carried out at high temperature or in the presence of an organic solvent such as formamide; these conditions “melt out” the short hairpin helices formed where base-pairing interactions occur within a single strand that folds back on itself.
- Bacterial cells could not survive such harsh conditions and instead use a single-strand binding protein, the SSB protein, to open their helices.
- This protein is essential for DNA replication as well as for general recombination in E. coli; it binds tightly and cooperatively to the sugar-phosphate backbone of all single-stranded regions of DNA, holding them in an extended conformation with their bases exposed.
- In this extended conformation a DNA single strand can base-pair efficiently with either a nucleoside triphosphate molecule (in DNA replication) or a complementary section of another DNA single strand (in genetic recombination).
- When hybridization reactions are carried out in vitro under conditions that mimic those inside a cell, the SSB protein speeds up the rate of DNA helix nucleation and thereby the overall rate of strand annealing by a factor of more than 1000.
General Genetic Recombination Usually Involves a Cross-Strand Exchange
- Exchanging a single strand between two double helices is presumed to be the slow and difficult step in a general recombination event (see Figure 3.8).
- After this initial exchange, extending the region of pairing and establishing further strand exchanges between the two closely apposed helices is thought to proceed rapidly.
- During these events a limited amount of nucleotide excision and local DNA resynthesis often occurs, resembling some of the events in DNA repair.
- Because of the large number of possibilities, different organisms are likely to follow different pathways at this stage.
- In most cases, however, an important intermediate structure, the cross-strand exchange, will be formed by the two participating DNA helices.
- One of the simplest ways in which this structure can form is shown in Figure 3.14.

Fig 3.14
- In the cross-strand exchange (also called a Holliday junction) the two homologous DNA helices that initially paired are held together by mutual exchange of two of the four strands present, one originating from each of the helices.
- No disruption of base-pairing is necessary to maintain this structure, which has two important properties
- (1) the point of exchange between the two homologous DNA double helices (where the two strands cross in Figure 3.14) can migrate rapidly back and forth along the helices by a double branch migration;
- (2) the cross-strand exchange contains two pairs of strands: one pair of crossing strands and one pair of non-crossing strands.
- The structure can isomerize, however, by undergoing a series of rotational movements, so that the two original non-crossing strands become crossing strands and vice versa (Figure 3.15).

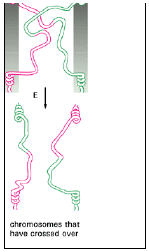
fig 3.15
- In order to regenerate two separate DNA helices and thus terminate the pairing process, the two crossing strands must be cut.
- If the crossing strands are cut before isomerization, the two original DNA helices separate from each other nearly unaltered, with only a very short piece of single stranded DNA exchanged.
- If the crossing strands are cut after isomerization, however, one section of each original DNA helix is joined to a section of the other DNA helix; in other words, the two DNA helices have crossed over (see Figure 3.15).
- The isomerization of the cross-strand exchange should occur spontaneously at some rate, but it may also be enzymatically driven or otherwise regulated by cells.
- Some kind of control probably operates during meiosis, when the two DNA double helices that pair are constrained in an elaborate structure called the synaptonemal complex
Gene Conversion Results from Combining General Recombination and Limited DNA Synthesis
- It is a fundamental law of genetics that each parent makes an equal genetic contribution to the offspring, one complete set of genes being inherited from the father and one from the mother.
- Thus, when a diploid cell undergoes meiosis to produce four haploid cells, exactly half of the genes in these cells should be maternal (genes that the diploid cell inherited from its mother) and the other half paternal (genes that the diploid cell inherited from its father).
- In a complex animal, such as a human, it is not possible to check this prediction directly.
- But in other organisms, such as fungi, where it is possible to recover and analyze all four of the daughter cells produced from a single cell by meiosis, one finds cases in which the standard genetic rules have apparently been violated.
- Occasionally, for example, meiosis yields three copies of the maternal version of a gene (allele) and only one copy of the paternal allele, indicating that one of the two copies of the paternal allele has been changed to a copy of the maternal allele.
- This phenomenon is known as gene conversion.
- It often occurs in association with general genetic recombination events, and it is thought to be important in the evolution of certain genes.
- Gene conversion is believed to be a straightforward consequence of the mechanisms of general recombination and DNA repair.
- During meiosis heteroduplex joints are formed at the sites of crossing-over between homologous maternal and paternal chromosomes.
- If the maternal and paternal DNA sequences are slightly different, the heteroduplex joint may include some mismatched base pairs.
- The resulting mismatch in the double helix may then be corrected by the DNA repair machinery, which either can erase nucleotides on the paternal strand and replace them with nucleotides that match the maternal strand or vice versa.
- The consequence of this mismatch repair will be a gene conversion.
- Gene conversion can also take place by a number of other mechanisms, but they all require some type of general recombination event that brings two copies of a closely related DNA sequence together.
- Because an extra copy of one of the two DNA sequences is generated, a limited amount of DNA synthesis must also be involved.
- Genetic studies show that usually only small sections of DNA undergo gene conversion, and in many cases only part of a gene is changed.
- Gene conversion can also occur in mitotic cells, but it does so more rarely.
- As in meiotic cells, some gene conversions in mitotic cells probably result from a mismatch repair process operating on heteroduplex DNA.
- Another likely mechanism in both meiotic and mitotic cells is illustrated in Figure 3.16.

fig 3.16
Mismatch Proofreading Can Prevent Promiscuous Genetic Recombination Between Two Poorly Matched DNA Sequences
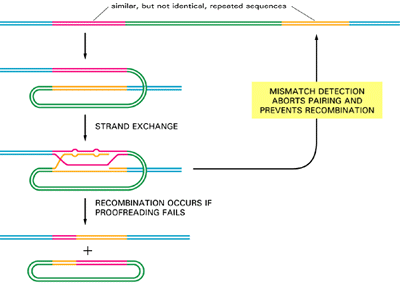
- As previously discussed, general recombination is triggered whenever two DNA strands of complementary sequence pair to form a heteroduplex joint between two double helices (see Figure 3.14).
- Experiments carried out in vitrowith purified RecA protein show that pairing can occur efficiently even when the sequences of the two DNA strands do not match well – when, for example, only four out of every five nucleotides on average can form base pairs.
- How, then, do vertebrate cells avoid promiscuous general recombination between the many thousands of copies of closely related DNA sequences that are repeated in their genomes?
- Although the answer is not known, studies with bacteria and yeasts demonstrate that the same mismatch proofreading system that removes replication errors has the additional role of interrupting genetic recombination events between imperfectly matched DNA sequences. It has long been known, for example, that homologous genes in two closely related bacteria,
- Escherichia coli and Salmonella typhimurium, generally will not recombine, even though their nucleotide sequences are 80% identical; when the mismatch proofreading system is inactivated by mutation, however, there is a 1000-fold increase in the frequency of such interspecies recombination events.
- It is thought, then, that the mismatch proofreading system normally recognizes the mispaired bases in an initial strand exchange and prevents the subsequent steps required to break and rejoin the two paired DNA helices.
- This mechanism protects the bacterial genome from the sequence changes thatwould otherwise be caused by recombination with foreign DNA molecules that occasionally enter the cell.
- In vertebrate cells, which contain many closely related DNA sequences, the same type of proofreading is thought to help prevent promiscuous recombin-ation events that would otherwise scramble the genome (Figure 3.17)