Physical Mapping of Genes on Chromosomes
- “Gene mapping” refers to the mapping of genes to specific locations on chromosomes. It is a critical step in the understanding of genetic diseases.
- There are two types of gene mapping:
Genetic Mapping –
- using linkage analysis to determine the relative position between two genes on a chromosome
Physical Mapping –
- using all available techniques or information to determine the absolute position of a gene on a chromosome.
- The ultimate goal of gene mapping is to clone genes, especially disease genes.
- Once a gene is cloned, we can determine its DNA sequence and study its protein product.
- For example, cystic fibrosis (CF) is the most common lethal inherited disease in the United States.
- As many as 1 in 2500 Americans of Northern European descent carries a gene with CF. In 1985, the gene was mapped to chromosome 7q31-q32 by linkage analysis.
- Four years later, it was cloned by Francis Collins and his co-workers.
- We now know that the disease is caused by the defect of a chloride channel – the protein product of this disease gene.
- Physical mapping techniques which are most common are following two:-
- Somatic cell hybridization and
- Fluorescent In situ Hybridization (FISH)
The resolution of FISH is much higher than somatic cell hybridization.
Mapping by DNA Sequencing
- Another means of physically mapping genes is to determine the sequence of nucleotides in the DNA.
- With this technique, physical distances between genes are measured in numbers of base pairs.
- Continuous sequences can be determined for only relatively small fragments of DNA; so, after sequencing, some method is still required to map the individual fragments.
- This mapping is often done by using the traditional gene mapping that examines rates of crossing over between molecular markers located on the fragments.
- It can also be accomplished by generating a set of overlapping fragments, sequencing each fragment, and then aligning the fragments by using a computer program that identifies the overlap in the sequence of adjacent fragments.
- With these methods, complete physical maps of entire genomes have been produced.
Physical Chromosome Mapping
- Genetic maps reveal the relative positions of genes on a chromosome on the basis of frequencies of crossing over, but they do not provide information that can allow us to place groups of linked genes on particular chromosomes.
- Furthermore, the units of a genetic map do not always precisely correspond to physical distances on the chromosome, because a number of factors other than physical distances between genes (such as the type and sex of the organism) can influence rates of crossing over.
- Because of these limitations, physical-mapping methods that do not rely on rates of crossing over have been developed.
Deletion Mapping
- One method for determining the chromosomal location of a gene is deletion mapping. Special staining methods have been developed that make it possible to detect chromosome deletions, mutations in which a part of a chromosome is missing.
- Genes are assigned to regions of particular chromosomes by studying the association of a gene’s phenotype or product and particular chromosome deletions.
- In deletion mapping, an individual that is homozygous for a recessive mutation in the gene of interest is crossed with an individual that is heterozygous for a deletion (Figure 5.15).
- If the gene of interest is in the region of the chromosome represented by the deletion (the red part of chromosome in Figure 5.15), approximately half of the progeny will display the mutant phenotype (see Figure 5.15a).
- If the gene is not within the deleted region, all of the progeny will be wild type (see Figure 5.15b).
- Deletion mapping has been used to reveal the chromosomal locations of a number of human genes.
- For example, Duchenne Muscular Dystrophy is a disease that causes progressive weakening and degeneration of the muscles.
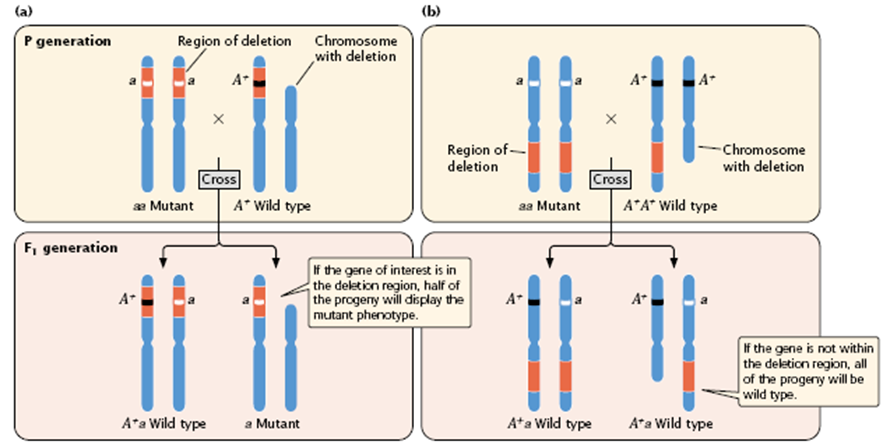
Figure 5.15 :
- Deletion mapping can be used to determine the chromosomal location of a gene. An individual homozygous for a recessive mutation in the gene of interest (aa) is crossed with an individual heterozygous for a deletion
- From its X-linked pattern of inheritance, the mutated allele causing this disorder was known to be on the X chromosome, but its precise location was uncertain.
- Examination of a number of patients having Duchenne Muscular Dystrophy, who also possessed small deletions, allowed researchers to position the gene to a small segment of the short arm of the X chromosome.
Somatic-Cell Hybridization
- Another method used for positioning genes on chromosomes is somatic cell hybridization, which requires the fusion of different types of cells.
- Most mature somatic (non sex) cells can undergo only a limited number of divisions and therefore cannot be grown continuously.
- However, cells that have been altered by viruses or derived from tumors that have lost the normal constraintson cell division will divide indefinitely; these types of cells can be cultured in the laboratory and are referred to as a cell line.
- Cells from two different cell lines can be fused by treating them with polyethylene glycol or other agents that alter their plasma membranes.
- After fusion, the cell possesses two nuclei and is called a heterokaryon.
- The two nuclei of a heterokaryon eventually also fuse, generating a hybrid cell that contains chromosomes from both cell lines.
- If human and mouse cells are mixed in the presence of polyethylene glycol, fusion results in human–mouse somatic-cell hybrids (Figure 5.16).

Figure 5.16 : Somatic-cell hybridization can be used to
- The hybrid cells tend to lose chromosomes as they divide and, for reasons that are not understood, chromosomes from one of the species are lost preferentially.
- In human–mouse somatic-cell hybrids, the human chromosomes tend to be lost, whereas the mouse chromosomes are retained.
- Eventually, the chromosome number stabilizes when all but a few of the human chromosomes have been lost.
- Chromosome loss is random and differs among cell lines.
- The presence of these extra” human chromosomes in the mouse genome makes it possible to assign human genes to specific chromosomes.
- n the first step of this procedure, hybrid cells must be separated from original parental cells that have not undergone hybridization.
- This separation is accomplished by using a selection method that allows hybrid cells to grow while suppressing the growth of parental cells.
- The most commonly used method is called HAT selection (Figure 5.17), which stands for hypoxanthine, aminopterin, and thymidine, three chemicals that are used to select for hybrid cells.

Figure 5.17 : HAT medium can be used to separate human–mouse hybrid cells from the original hybridized cells.
- In the presence of HAT medium, a cell must possess two enzymes to synthesize DNA: thymidine kinase (TK) and hypoxanthine-guanine phosphoribosyl transferase (HPRT).
- Cells that are tk – or hprt – cannot synthesize DNA and will not grow on HAT medium. The mouse cells used in the hybridization procedure are deficient in TK, but can produce HPRT (the cells are tk – hprt +); the human cells can produce TK but are deficient for HPRT (they are tk + hprt – ).
- On HAT medium, the mouse cells do not survive, because they are tk – ; the human cells do not survive, because they are hprt – .
- Hybrid cells, on the other hand, inherit the ability to make HPRT from the mouse cell and the ability to make TK from the human cell; thus, they produce both enzymes (the cells are tk + hprt +) and will grow on HAT medium.

Figure 5.18 : Somatic-cell hybridization is used to assign a gene to a particular human chromosome.
- A panel of six cell lines, each line containing a different subset of human chromosomes, is examined for the presence of the gene product (such as an enzyme).
- A plus sign means that the gene product is present; a minus sign means that the gene product is missing.
- Four of the cell lines (A, B, D, and F) have the gene product, indicating that the gene is present on one of the chromosomes found in these cell lines.
- The only chromosome common to all four of these cell lines is chromosome 4, indicating that the gene is located on this chromosome.
- To map genes using somatic-cell hybridization requires the use of a panel of different hybrid cell lines.
- The cell lines of the panel differ in the human chromosomes that they have retained.
- For example, one cell line might possess human chromosomes 2, 4, 7, and 8, whereas another might possess chromosomes 4, 19, and 20.
- Each cell line in the panel is examined for evidence of a particular human gene
- The human gene can be detected either by looking for the protein that it produces or by looking for the gene itself with the use of molecular probes.
- Correlation of the presence of the gene with the presence of specific human chromosomes often allows the gene to be assigned to the correct chromosome.
- For example, if a gene was detected in both of the aforementioned cell lines, the gene must be on chromosome 4, because it is the only human chromosome common to both cell lines (Figure 5.18).
- Two genes determined to be on the same chromosome with the use of somatic-cell hybridization are said to be syntenicgenes.
- This term is usedbecause syntenic genes may or may not exhibit linkage in the traditional genetic sense-remember that two genes can be located on the same chromosome but may be so far apart that they assort independently.
- Synthenic refers to genes that are physically linked, regardless of whether they exhibit genetic linkage. (Synteny is sometimes also used to refer to gene loci in different organisms located on a chromosome region of common evolutionary origin.)
- Sometimes somatic-cell hybridization can be used to position a gene on a specific part of a chromosome.
- Some hybrid cell lines carry a human chromosome with a chromosome mutation such as a deletion or a translocation.
- If the gene is present in a cell line with the intact chromosome but missing from a line with a chromosome deletion, the gene must be located in the deleted region (Figure 5.19) Similarly, if a gene is usually absent from a chromosome but consistently appears whenever a translocation (a piece of another chromosome that has broken off and attached itself to the chromosome in question) is present, it must be present on the translocated part of the chromosome.

Figure 5.19: 7.21 Genes can be localized to a specific part of a chromosome by using somatic-cell hybridization
- Note: If the gene product is present in a cell line with an intact chromosome but missing from a line with a chromosome deletion, the gene for that product must be located in the deleted region.






