ANEUPLOIDY
- Aneuploidy can be either due to loss of any one or more chromosomes to complete chromosome set (Hypoploidy).
- Hypoploidy is due to loss of single chromosome is known as monosomy, and the individual that carries such number is known as monosomics.
- The formula for monosomy is 2n-1. Another type of hypo aneuploidy is nullisomy – where the loss of a pair chromosome from diploid set. Both chromosome of a diploid set is absent.
- The general formula for nullisomy is 2n-(1×2) or 2n-2.
Origin of aneuploidy
Monosomy –
- When monosomy is present in diploid organism it can not be tolerated because one complete chromosome is lacking.
- Numbers of genes present in this chromosome were affected.
- This unbalancing creates a problem, but when monosomy is present in polyploid organism these could be easily produced and tolerable because more than two chromosome of same combination is present.
- Meiosis in monosomics behaves like haploids.
- In monosomics one of the chromosomes will go to only one pole during division and other pole will be devoid of it.
- Other chromosomes will asset normally. In other words, monosomics behave like a haploid for one chromosome and as a diploid for others.
Nullisomy –
- Another type of hypo-aneuploidy where a loss of pair of homologous chromosome is seen.
- When it found in diploid organism the organism doesn’t tolerate.
- The gametes produced by these individual are devoid of one chromosome – means they are aneuploids.
- The frequency of sterility is more in animals than in plants because the plants are able to tolerate loses of chromosomes much better than animals.
MONOSOMICS
- Numbers of chromosomes in Nicotiana tabacum are 24, and all the 24 monosomics for that plant were reported by Clausenand Camerson (1944).
- Same work was done by Sears (1954) for hexaploid bread wheat.
- A set of monosomic for Gossypium hirsutum was reported by Endrizzi.
Methods of production of monosomics
- From haploids – Sears (1939) got accidentally two haploids when he crossed two Geneva – (Secale cereale and Chinese springwheat).
- That haploid is used as female parent and crossed with normal hexaploid male.
- The seeds produced by this method showing monosomics.
- When these monosomics are selfed (inter crossing) nullisomic, trisomic and then later on tetrasomics were found by selfing.
- From backcrosses of inter specific hybrids – In N. tabacum monosomics have been produced by crossing the polyploidy species with one of its progenitors.
- The resultant F1 plant is backcrossed with polyploidy crop by which monosomics are found.
- Eg: In this eg. The haploid (n = 24) N. tabacum having 12 chromosomes of N. Sylvestris (n = 12) and 12 chromosomes of N. tomentosa (n = 12).
- This crop i.e. N. tabacum is crossed with N. sylvestris or (n = 24) develops F1 plant. Obtained F1 plants has 12 pairs chromosomes of N. sylvestris species and 12 single chromosomes of N. tomentosa.
- The gametes of this crop contain n = 12 to 24 as they have 21 chromosomes from N. sylvestris but the no. of single 12 chromosomes of N. tomentosa varies from gametes to gametes resulting in n = 12 to 24.
- These are then backcrossed with N. tabacum (n = 24) to develop monosomics for N. tomentosa.

Fig.4.2 Production of monosomics through interspecific hybridization in tobacco.
- From partially asynaptic plants – In this type during meiotic metaphase I, perfect bivalent formation is not observed, a variable number of univalents are observed.
- These univalents will be distributed to the two poles randomly during anaphase I.
- Irradiation treatment – In case of Gossypium hirsutum (cotton) and Avena sativa (Oats), irradiation of inflorescence to the production of gametes with n+1 or n-1. It is a type of artificial treatment.
- Spontaneous production – Monosomics have been observed in wheat, tobacco, cotton by natural occasional non-disjunction of a bivalent during meiosis.
Description and Identification of monosomics
- Sometimes monosomics are morphologically different or sometimes it may not. It depends on genes present on that particular chromosome for which plant is monosomic.
- Monosomic condition is confirmed through chromosome count during mitotic metaphase and by the use of a univalent at meiosis.
Meiotic behaviour of monosomics
- Monosomics usually form bivalents in addition to a solitary univalent at end of prophase I and metaphase I.
Transmission of monosomics
- Breeding behaviour of monosomics has been studied by examining the progeny obtained on selfing them and on crossing them separately as female and male parents with the normal.
- On the basis of these studies, it is always possible to calculate the frequency of functional deficient (n = 2x-1 or 3x-1) gametes relative to normal (n = 2x or 3x) type of gametes.
- Although deficient gametes are produced at a frequency higher than 75%, but they function in the pollen at a very low frequency in wheat (4%), but at a variable frequency in oats (1%-74%).
- In tobacco, although deficient pollen may function at a low frequency, nullisomic zygotes, which normally do not survive.
- This situation leads to the production of nullisomics in the progenies of selfed monosomics, in variable frequency.
Eg, In case of wheat monosomics, (where normally n = 3x = 21), functional male gametes are of 2 types,
i) n = 3x = 21, ii) n = 3x-1 = 20.
- The first category, male gametes are 96% while in second case only 4% gametes are produced.
- But in female gametes 25% gametes carrying n = 3x = 21 while remaining 75% having n = 3x-1 = 20.
- When these types of gametes crossed, percentage of nullisomics progeny is 3%, monosomics progeny is 73% and normal progeny is 24%.
TRISOMICS
Trisomics in Diploids
- The first aneuploid chromosome number was reported in Oenothera hybrids by Lutz (1909) and by Gates (1909) who recorded n = 15 while normal has n = 14.
Trisomy –
- Trisomics is an example of hyperaneuploidy where an organism has an extra chromosome (2n+1).
- Since the extra chromosome may belong to anyone of the different chromosomes of a haploid complement.
- The number of possible trisomics in an organism will be equal to its haploid chromosome number.
- Eg, Barley the haploid chromosome no. is 7 (n = 7) consequently, 7 trisomics are possible.
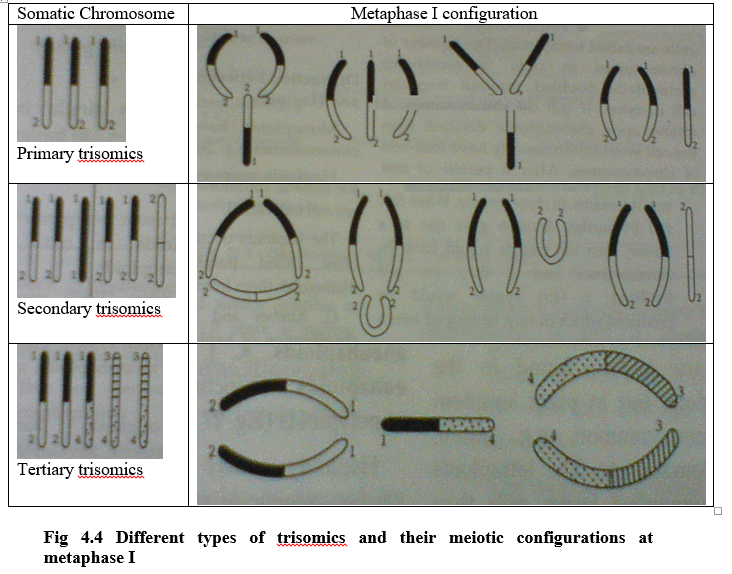
Trisomics are classified into 3 categories
- Primary Trisomics – where an extra chromosome is identical to two homologous chromosomes (diploid) in their gene sequence.
- Secondary Trisomics – where the extra chromosome having same gene in multiple copies (i.e. isochromosome). The extra chromosome having duplicated and deficient gene.
- Tertiary Trisomics – The extra chromosome should be product of translocation i.e. this chromosome having genes of 2 or more other chromosomes.
- Trisomics were obtained for the firsttime in Datura stramonium by A. F. Blakeslee and his coworkers.
- The genomic number of this species is n = 12. Blakeslee and his coworkers found 12 primary trisomics, 24 secondary trisomics and large no. of possible tertiary trisomics.
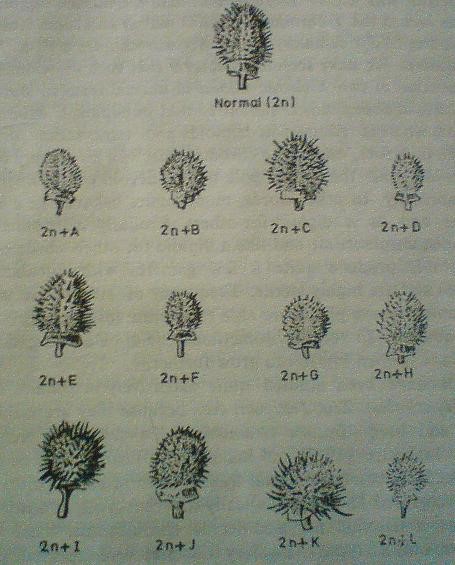
- In Datura stramonium most of the trisomic were identified by the morphological features of fruit by shape and size.


Origin and source of trisomics
- Trisomics may originate spontaneously due to production of (n+1) type of gametes; this gamete develops due to non-disjunction of a bivalent.
- Mostly trisomics are produced by artificial methods.
Cytology of trisomics
- A trisomic has an extra chromosome, which is homologous to one of the chromosomes of the complement.
- During meiosis all of them synapse with each other and at metaphase I they arrange themselves from a peculiar shape of arrangement. (Fig 4.4)
Use of monosomics and trisomics in chromosomal mapping of diploid and polyploidy species
- The result of trisomic ratios can be utilized for locating genes on specific chromosome or for finding out distances of these genes from centromere, this technique is called chromosome mapping.
- If linkage groups are already established in an organism, trisomics can be effectively used for assigning these linkage groups to specific chromosomes.
- Since the segregation ratio for genes located on the chromosome involved in trisomic condition differs from that, genes located on other chromosomes, through these ratios one can found out on which chromosome a particular gene is located.
- This technique has been successfully applied in maize, tomato, barley, datura and Arabidopsis.



